Harvard Children's Hospital Informatics Program
Lisp and SNPer help biologists speed up the data analysis process
 The massive amount of data generated via the mapping of the human genome has given scientists a tremendous edge in understanding disease ֠but it is overwhelming as well. Now, biologists must move beyond simply identifying the data, and figure out how to use it to find solutions. Lisp and Allegro CL are helping them meet this challenge.
The massive amount of data generated via the mapping of the human genome has given scientists a tremendous edge in understanding disease ֠but it is overwhelming as well. Now, biologists must move beyond simply identifying the data, and figure out how to use it to find solutions. Lisp and Allegro CL are helping them meet this challenge.
The Children's Hospital Informatics Program (http://chip.org/) is a multidisciplinary applied research and education program at Children's Hospital in Boston, focusing on Bioinformatics and Clinical Informatics. Faculty members are involved in a large number of research activities ranging from microarray data analysis, to the development of Bioinformatics software and databases, to the exploration of the relationships between genetic and clinical data.
One of their projects, SNPper, is built in Allegro CL and Lisp, and provides scientists with a variety of tools that greatly speed up the SNP (Single Nucleotide Polymorphisms) analysis process. SNPs are mutations that affect a single base (A, T, G or C) in our DNA. They are responsible not only for most of the differences between two human beings, but also for a variety of diseases, medical conditions, altered response to drugs, etc. There are over 3 million currently known SNPs
Using SNPper, scientists can search for SNPs in public genomic databases, retrieve sets of them according to their position on the genome, filter them according to the desired characteristics, and export the corresponding data in a variety of formats (HTML tables, structured XML documents, images, etc). Users of the system can now accomplish the same amount of work in a day that used to take weeks to complete.
Dr. Alberto Riva, the Project Leader for SNPper, began developing the application after seeing a biologist spend days cutting and pasting data across three different databases. He used Lisp and Allegro CL because he needed a language and platform able to deal with huge data structures, increased knowledge exchange needs and under-specified and evolving analysis methods.
"Lisp is the only language that allows me to concentrate on solving interesting problems rather than dealing with trivial syntax/memory/debugging issues." Riva says. "Common Lisp allows my programs to grow and evolve, instead of forcing me to constantly reinvent wheels. Its stability, performance, excellent debugging features, and extensive standard library are all essential for my work" he adds.
Riva also believes that Lisp is the language of choice for the computational biology tools of the future. He says that most Bioinformatics tools have been developed in an ad-hoc way, to solve specific problems in isolation. This has resulted in poor interoperability, lack of data standards, and limited integration.
"Today, we need to deal with complete genomes composed of billions of bases, microarray experiments that generate thousands of data points, databases holding data on tens of thousands of proteins. The next generation of computational biology tools will therefore require a new approach. Lisp is the ideal language to support the evolution of our Bioinformatics in this direction." He emphasizes.
SNPper is built using Allegro CL 6.2 on GNU/Linux (RedHat 7.1). The underlying database is MySQL. For more information about SNPper, please visit http://snpper.chip.org/.
Click here to download a PDF version of this story.
| Copyright © 2023 Franz Inc., All Rights Reserved | Privacy Statement | 
|

|

|

|
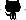
|








