Introduction
The file Neuro-Symbolic AI and Large Language Models Introduction introduces support for Neuro-Symbolic AI and Large Language Models in Allegro CL 8.0.0. In this file we provide multiple examples using these tools. Almost all can be copied and pasted to the New WebView query page.
Note examples with an AI key use a fake key that will not work. You have to replace that with your actual key. See the section You need an API key to use any of the LLM predicates in llm.html for information on obtaining a key.
Getting started
Start AllegroGraph 8.0.0 and display the New WebView page (the traditional WebView page does not have an LLM Playground link). Do this by opening AllegroGraph is a browser and clicking on the New WebView choice on the menubar just below the banner.
The page look like:
Click on the Create LLM Playground button. This will display a dialog asking for an OpenAI API Key (the illustration shows an invalid key which has the format of a valid one, sk- followed by numbers and letters):
Once you supply your key and it is accepted a repo named llm-playground-1 will be created (and subsequent repos will be name llm-playground-2, -3, and so on). You are now in an environment where LLM-related queries can be posed.
The repo has a number of stored queries relevant to LLM, has the llm: namespace defined (as http://franz.com/ns/allegrograph/8.0.0/llm/ which is the prefix for LLM magic properties) and the query option openaiApiKey is set up with the OpenAI Key you supplied. You can set up any repo to have these features (you have to add and save the stored queries if you want them, just copy and paste from the LLM playground repo). We show how to do that in the Embedding example below.
Use of Magic predicates and functions
AllegroGraph has a number of predefined magic predicates and functions which can be used in LLM-related queries. The magic predicates are listed in the SPARQL Magic Properties document. A magic predicate can be used in the predicate position in queries and functions can transform the value of a variable. These significantly extend the power of SPARQL queries. There are examples below which will make this clear.
Specifically, we introduce three magic predicates, two magic functions, and a command line option for agtool. The combination of these simple primitives provides direct access from SPARQL to Large Language Model responses and Vector Store document matching.
The supported operations are (see here for magic functions and here for magic predicates):
llm:response: Both a function and a predicate, depending on whether we want the LLM to return one or a list of items.- llm:node: A function to generate a unique URI for a text literal.
- llm:nearestNeighbor: A predicate that works on AllegroGraph Vector store. It takes as input a string or query and finds the best matches in the Vector Store.
- llm:askMyDocuments: A high level predicate to ask the LLM about information in our local Vector Store.
- llm:askForTable: A high level predicate to ask the LLM about information and return results in tabular form.
The llm: is a predefined prefix which expands to
<http://franz.com/ns/allegrograph/8.0.0/llm/> We supply this prefix in every query but you can easily add it to the list of defined namespaces as described in Namespaces and query options.
Power of LLM
Large Language Models provide a fountain of data of interest to anyone concerned with Graph Databases. AllegroGraph can now retrieve responses from Large Language Models, utilize those responses to
- Generate tabular data
- Build custom knowledge graphs
- Curate information in existing databases
- Match words, concepts, sentences, and queries with embeddings
- Perform retrieval-augmented generation
In this tutorial, we will explore how to use magic predicates and functions to connect the knowledge graph to Large Language Models.
Pre-defined queries
The LLM Playground contains 7 Saved Queries you can experiment with. Let’s review them one by one:
Query 0 - Hello, GPT
# This query demonstrates the use of the `response` SPARQL function.
# It returns a single string response to the given prompt.
PREFIX llm: <http://franz.com/ns/allegrograph/8.0.0/llm/>
SELECT ?response {
BIND (llm:response("Hello, GPT.") AS ?response)
} The magic function llm:response sends its argument to the LLM and binds its response to ?response. Because all natural language is text, the value bound to ?response is a literal string.
You can experiment with replacing the input "Hello, GPT" with your own prompts.
Query 1 - Entry Lists
llm:response is the name of both a function and a magic predicate. We use the function when we want the LLM to return a single value. When we want to bind multiple values, we use the llm:response magic predicate.
This query demonstrates the use of response magic predicate. It generates a list of responses bound to the SPARQL variable ?entry.
PREFIX llm: <http://franz.com/ns/allegrograph/8.0.0/llm/>
SELECT ?entry {
?entry llm:response "List the US States.".
# Also try:
# ?entry llm:response "Suggest some names for a male cat."
# ?entry llm:response "List causes of the American Revolutionary War."
# ?entry llm:response "Name the colors of the rainbow."
# ?entry llm:response "List some species of oak.
} Try some of the prompts in the commented examples and also try some of your own. There is no restriction on the prompt except that you want the prompt to request a list or array of items.
Query 2 - Resources
When building datasets, we need to be able to turn string responses into IRIs which will be used as subjects in statements. In order to make an IRI out of a response string, use function node. It converts a string into a unique fixed-size IRI in a deterministic way.
PREFIX llm: <http://franz.com/ns/allegrograph/8.0.0/llm/>
SELECT ?state ?node {
BIND (llm:response("Largest US state, name only.") AS ?state)
BIND (llm:node(?state) AS ?node)
# Node can also be optionally returned by a magic property:
# (?state ?node) llm:response "List US states.".
} The result is:
state node
"Alaska" llm:node#7a067bb6f837b5fd800a16dbbd18c629
This query introduces a function llm:node that creates a unique IRI for a given response string. This IRI is useful when we want to create linked data. The LLM returns only literal strings, which may be useful as node labels, but often we’d like to have a unique IRI for each LLM item generated.
The commented query illustrates the use of the optional ?node output variable in llm:response. The ?node is bound to a unique IRI for each of the values bound to ?state.
Query 3 - States
This query demonstrates the combination of both the response magic property and the response function to produce a table of US States and some properties of those states: capital, population, area, year of admission, governor and a famous celebrity from that state. Beware that the query may take up to 60 seconds to run: it makes 61 API calls to the OpenAI API. We limited the number of states in the subquery to 10 in order to reduce the response time.
PREFIX llm: <http://franz.com/ns/allegrograph/8.0.0/llm/>
SELECT ?state ?capital ?pop ?area ?admit ?gov ?celeb {
{
SELECT ?state { ?state llm:response "List the US states.". }
ORDER BY RAND()
LIMIT 10
}
BIND (llm:response(CONCAT("Name the capital of the state of ", ?state, ". Return the city name only.")) AS ?capital).
BIND (llm:response(CONCAT("State the population of the state of ", ?state, ". Respond with a number only.")) AS ?pop).
BIND (llm:response(CONCAT("Tell me the square mile area of the state of ", ?state, ". Respond with a number only.")) AS ?area).
BIND (llm:response(CONCAT("In what year was ", ?state, " admitted to the Union?. Respond with a year only.")) AS ?admit).
BIND (llm:response(CONCAT("Who is the governor of the state of ", ?state, "?. Respond with the governor's name only.")) AS ?gov).
BIND (llm:response(CONCAT("Who is the most famous celebrity from the state of ", ?state, "?. Respond with the celebrity's name only.")) AS ?celeb).
}
ORDER BY ?state In this example we generate a table of data utilizing the llm:response predicate in conjunction with the llm:response function. The predicate returns a list of states and the function returns the value of a distinct property (capital, population, etc.)
You may notice when running this query that it seems to take a long time. This is because each call to llm:response requires an API call to OpenAI, and the server spends most of time waiting for the API to return.
Here is part of the result:
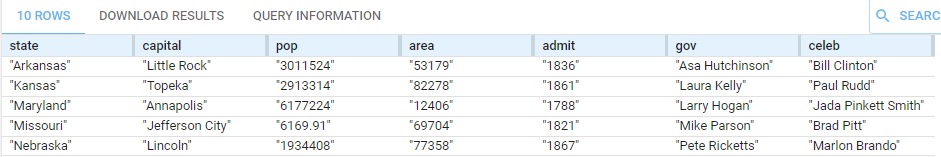
See also the askForTable states example which is similar but the query is shorter and uses different tools.
Query 4 - Borders
The previous query generated a table of data, much like the result of a SQL query on a relational database. The true power of LLMs in the graph database comes into focus when we generate the relationship between entities.
This query demonstrates using GPT to generate linked data. It populates a database with the border relationships between states. The query utilizes the optional second subject argument ?node in response magic predicate. The ?node variable is bound to a unique IRI which is deterministically computed from the response string. A validation step at the end deletes any bordering relationships that are not symmetric. Because we're executing INSERT and DELETE, "No table results" will appear after execution. Beware that the query may take up to 50 seconds to run: it makes 51 API calls to the OpenAI API.
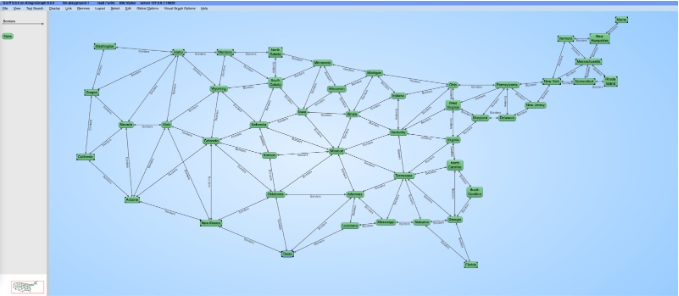
Once this query finishes, you can visualize the graph of border relationships between states. In order to do that, open Gruff and copy the query here into the Query View. You should see a topologically accurate map of the continental United States (this portion is commented out because it is for Gruff, the query starts with the PREFIX).
#
# SELECT ?state1 ?state2 { ?state1 <http://franz.com/borders> ?state2 }
#
PREFIX llm: <http://franz.com/ns/allegrograph/8.0.0/llm/>
PREFIX : <http://franz.com/>
CLEAR ALL;
INSERT {
?stateNode rdfs:label ?state.
?stateNode rdf:type :State.
?stateNode :borders ?bordersNode.
} WHERE {
(?state ?stateNode) llm:response "List the US States.".
BIND (CONCAT("Write the states that border this state: ", STR(?state)) AS ?bordersPrompt).
(?borders ?bordersNode) llm:response ?bordersPrompt.
};
DELETE {
?state1 :borders ?state2
} WHERE {
?state1 :borders ?state2.
FILTER NOT EXISTS { ?state2 :borders ?state1 }
}
The Borders query differs from the previous examples in several ways:
We want to
INSERTdata into the graph. Running the query by itself displays no results.The
DELETEstatement serves as a validation step. Sometimes the LLM hallucinates a bordering relation when none exists, but generally will not “double-hallucinate” a border in both directions. So theDELETEremoves and bordering relations where there is evidence that X borders Y but none that Y borders X.
We’ll use Gruff to visualize the results:
Query 5 - Ontology
This query demonstrates how we can generate an ontology out of thin air, even for the most obscure topics. We're asking GPT to generate a taxonomy of the the Areas of Life. At each level, we've asked GPT to limit the branching factor, but it's easy to see that as the depth of the taxonomy increases, and the branching factor widens, the time complexity of this query can grow rapidly. This example takes about 90 seconds to run for only 2 life areas and because it only INSERTs data, it returns "No table results".
Once this query finishes, you can visualize the Areas of Life Ontology. In order to do this, open Gruff and copy the query here into the Query View. Try selecting the <http://franz.com/AreaOfLife> node and choose the menu option "Layout -> Do Tree Layout from Selected Node".
#
# SELECT DISTINCT ?area ?concept ?habit ?trait {
# ?concept <http://franz.com/hasTrait> ?trait.
# ?trait <http://franz.com/hasHabit> ?habit.
# ?area rdf:type <http://franz.com/AreaOfLife>; <http://franz.com/hasConcept> ?concept.
# } LIMIT 32
#
# You can increase the number of results in Gruff by removing the limit
# or increasing the LIMIT value.
PREFIX llm: <http://franz.com/ns/allegrograph/8.0.0/llm/>
PREFIX : <http://franz.com/>
CLEAR ALL;
INSERT {
?areaUri rdf:type :AreaOfLife.
?conceptUri rdf:type :Concept.
?traitUri rdf:type :Trait.
?habitUri rdf:type :Habit.
?areaUri rdfs:label ?area.
?conceptUri rdfs:label ?concept.
?traitUri rdfs:label ?trait.
?habitUri rdfs:label ?habit.
?areaUri :hasConcept ?conceptUri.
?conceptUri :hasTrait ?traitUri.
?traitUri :hasHabit ?habitUri.
} WHERE {
# Comment out the VALUES clause and uncomment this subquery to see
# what areas GPT recommends:
# {
# SELECT ?area ?areaUri {
# (?area ?areaNode) llm:response "List the 10 general areas of life in the wheel of life.".
# }
# LIMIT 10
# }
# Uncomment the remaining areas in the VALUES clause to generate a
# more complete ontology.
VALUES ?area {
"Family"
"Recreation"
# "Self-image"
# "Health"
# "Career"
# "Contribution"
# "Spirituality"
# "Social"
# "Love"
# "Finance"
}
BIND (llm:node(?area) AS ?areaUri).
BIND (CONCAT("In the ", ?area, " area of life, list 5 to 10 important general concepts.") AS ?conceptPrompt).
(?concept ?conceptUri) llm:response ?conceptPrompt.
BIND (CONCAT("For the concept ", ?concept, " in the ", ?area, " area of life, list 2 to 4 traits or preferences a person might have or lack in his relationship with ", ?concept, ".") AS ?traitPrompt).
(?trait ?traitUri) llm:response ?traitPrompt.
BIND (CONCAT("List 2 to 4 habits or activities associated with ", ?concept, " trait ", ?trait, " in the ", ?area, " area of life") AS ?habitPrompt).
(?habit ?habitUri) llm:response ?habitPrompt.
} Like the previous query, this one uses INSERT to add new triples to the graph database. Also the query by itself displays no results: we can use Gruff to visualize the generated triples.
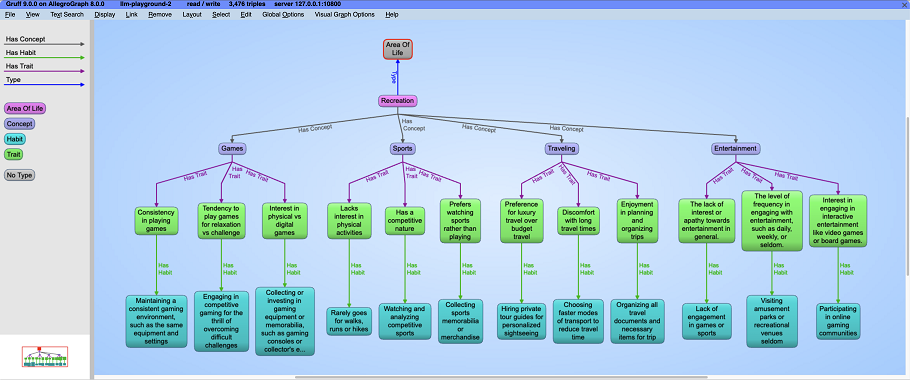
You can experiment with uncommenting some of the ?area values to generate a much larger ontology. You can also uncomment the inner SELECT clause to have GPT generate the areas of life for you. GPT may suggest a different organization of life areas. Beware that the more terms you include, the longer the query will run!
Query 6 - Summary
The final sample query in the LLM Playground illustrates document summarization.
This query is a simple example of text summarization. Typically we might want to summarize document text from pre-existing documents. But in this demo, we simply ask GPT to generate some long essays and then summarize them. This query may take around 80 seconds to run: it only does 9 OpenAI API calls, but 4 of them return sizeable responses (the essays).
PREFIX llm: <http://franz.com/ns/allegrograph/8.0.0/llm/>
SELECT ?topic ?essay ?summary {
?topic llm:response "List 4 essay topics."
BIND (llm:response(CONCAT("Write a long essay on the topic ", ?topic)) AS ?essay).
BIND (llm:response(CONCAT("Provide a very brief summary of this essay about ", ?topic, ": ", ?essay)) AS ?summary).
} Here is the very beginning of the results (just part of the first essay and the summary). Looking at the scrollbar to the right you can see how much output was generated.

Data Validation
Now let’s look at a practical example of using LLM magic predicates and functions in a real-world application. The Unified Medical Language System (UMLS) is a large repository of some 800,000 concepts from the biomedical domain, organized by several million interconcept relationships. UMLS is known to have many inconsistencies including circular hierarchical relationships. This means that one concept can be linked to itself through a circular path of narrower or broader relationships. Many of the circular relationships in UMLS are simple 1-cycles where two concepts are both set to be narrower than the other.
To declutter the knowledge graph, we delete relationships under certain conditions. Specifically we look for relations between concepts where X is narrower than Y, and Y is narrower than X. We then retrieve the labels of those concepts and build prompts for GPT asking GPT to define those concepts. Once we have those definitions we can build another prompt that says “based on these definitions, is the X concept really a narrower or a more specific concept than the Y?” and another prompt that asks the opposite, “Is Y really narrower than X?” We ask GPT to answer yes or no.
Here are some of the responses that GPT gave us about the narrower relationships. When GPT says “Yes” to “Is X narrower than Y” and “No” to “is Y narrower than X”, we can confidently delete the first relationship. When “No” to the first and Yes” to the second, we can delete the second. Only when GPT answers “Yes” or “No” to both questions are we left in doubt, and perhaps do something else such as delete both, or replace both with skos:exactMatch (if the answer to both questions is “Yes”).
xlabel: "Primary malignant neoplasm of extrahepatic bile duct"
ylabel: "Malignant tumor of extrahepatic bile duct"
xyResponse: "No"
yxResponse: "No"
xlabel: "Accident due to excessive cold"
ylabel: "Effects of low temperature"
xyResponse: "Yes"
yxResponse: "No"
xlabel: "Echography of kidney"
ylabel: "Ultrasonography of bilateral kidneys"
xyResponse: "No"
yxResponse: "Yes"
xlabel: "Skin of part of hand"
ylabel: "Skin structure of hand"
xyResponse: "No"
yxResponse: "Yes"
xlabel: "Removal of urinary system catheter"
ylabel: "Removal of bladder catheter"
xyResponse: "No"
yxResponse: "Yes"
xlabel: "Degloving injury of head and neck"
ylabel: "Degloving injury of neck"
xyResponse: "Yes"
yxResponse: "No"
xlabel: "Other noninfective gastroenteritis and colitis"
ylabel: "Noninfective gastroenteritis and colitis, unspecified"
xyResponse: "No"
yxResponse: "No"
xlabel: "Open wound of lower limb"
ylabel: "Multiple and unspecified open wound of lower limb"
xyResponse: "No"
yxResponse: "Yes"
xlabel: "Mandibular left central incisor"
ylabel: "Structure of permanent mandibular left central incisor tooth"
xyResponse: "No"
yxResponse: "Yes"
The full query shown here executes the DELETE when the FILTER statement is true.
PREFIX llm: <http://franz.com/ns/allegrograph/8.0.0/llm/>
DELETE {
?x skos:narrower ?y
} WHERE {
{SELECT ?x ?y {
?x skos:narrower ?y.
?y skos:narrower ?x.
} ORDER BY RAND() LIMIT 16}
?x skos:prefLabel ?xlabel.
?y skos:prefLabel ?ylabel.
bind(concat("Define ",?xlabel,": ") as ?xdefPrompt).
bind(concat("Define ",?ylabel,": ") as ?ydefPrompt).
bind( llm:response (?xdefPrompt) as ?xdef)
bind( llm:response (?ydefPrompt) as ?ydef)
bind(concat(
"Based on these definitions: ",
?xlabel,
": ",
?xdef,
" and ",
?ylabel,
": ",
?ydef,
", is ",
?xlabel,
" really a narrower, more specific concept than ",
?ylabel,
"? Answer yes or no.") as ?xyPrompt).
?xyResponse llm:gptSays ?xyPrompt.
bind(concat(
"Based on these definitions: ",
?xlabel,
": ",
?xdef,
" and ",
?ylabel,
": ",
?ydef,
", is ",
?ylabel,
" really a narrower, more specific concept than ",
?xlabel,
"? Answer yes or no.") as ?yxPrompt).
?yxResponse llm:gptSays ?yxPrompt.
FILTER(regex(ucase(?xyResponse), "^NO") && regex(ucase(?yxResponse), "^YES"))
} Note that we’ve implemented a subquery to select a random sample of 16 1-cycles. This is because the query is very slow: again, spending most of its time waiting for API responses. We might prefer to run this query in shell script using agtool:
% agtool query SERVER-SPEC/umls query.rq Where query.rq contains the SPARQL DELETE query above. Running this agtool command in a loop can eliminate about 200 1-cycles per hour. We found that the UMLS knowledge base contains 12690 1-cycles, so a full purge could take around 60 hours.
askForTable example
The askForTable magic predicate allows repsonses to be displayed in a table. Here are a couple of examples. Run them in the LLM playground repo created above.
PREFIX llm: <http://franz.com/ns/allegrograph/8.0.0/llm/>
SELECT ?language ?a ?b ?c ?d ?e ?f ?g ?h ?i ?j ?k ?l ?m ?n ?o ?p ?q ?r ?s ?u ?v ?w ?x ?y ?z {
(?language ?a ?b ?c ?d ?e ?f ?g ?h ?i ?j ?k ?l ?m ?n ?o ?p ?q ?r ?s ?u ?v ?w ?x ?y ?z ) llm:askForTable "Make a table of 27 columns, one for the language and one for each letter of the Roman Alphabet. Now, fill in each row with the nearest matching phonetic letters from the Greek, Cyrillic, Hebrew, Arabic, and Japanese languages."} 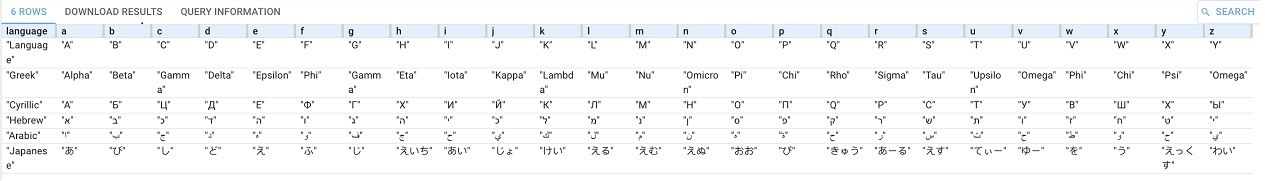
This example is similar to query 3 above but uses askForTable.
PREFIX llm: <http://franz.com/ns/allegrograph/8.0.0/llm/>
SELECT * {
(?state ?capital ?population ?area ?admit ?governor ?celebrity) llm:askForTable "List the US states, their capital, population, square mile area, year admitted to the Union, the current governor and the mst famous celebrity from that state."} 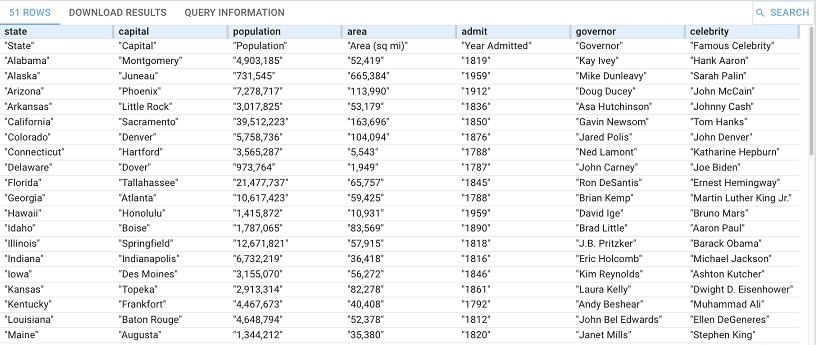
Embedding by creating a VectorStore
An embedding is a large-dimensional vector representing the meaning of some text. An LLM creates an embedding by tokenizing some text, associating a number with each token, and then feeds the numbers into a neural network to produce the embedding vector. The text can be a word (or even, a letter), a phrase, a sentence, or a longer body of text up to a length determined by token limit of the LLM. A really useful feature of embeddings is that semantically similar text maps to similar embeddings. “Fever, chills, body aches, fatigue” has a similar embedding vector to “flu-like symptoms.” Given a large database of embeddings, we can measure the distance between a target text and the existing embeddings, and determine the nearest neighbor matches of the target.
LLMs can compute embeddings of text up to their token limit size (8191 tokens for OpenAI text-embedding-ada-002, or about 6000 words). Many applications of embeddings in fact rely on very large text blocks. But for expository purposes, it’s easier to understand and demonstrate the applications using much shorter strings, so we will start with a collection of short text labels first.
For this example, we first create a new repo historicalFigures (here is the New WebView dialog creating the repo:
Next we have to associate our OpenAI key with the repo. Go to Repository | Repository Control | Query execution options (type "Query" into the search box after Repository control is displayed) and enter the label openaiApiKey and your key as the value, then click SAVE QUERY OPTIONS:
(The key in the picture is not valid.) You can also skip this and use this PREFIX with every query (using your valid key, of course):
PREFIX franzOption_openaiApiKey: <franz:sk-U01ABc2defGHIJKlmnOpQ3RstvVWxyZABcD4eFG5jiJKlmno> We will simply use the llm: prefix in every query we run. You can add it as a defined namespace if you wish to. Click on Namespaces in the Repository menu to the left and click on NEW NAMESPACE.
We create a very simple repository of historical figures using LLM. Note we define the llm: prefix. (These set-up steps, associating the openaiApiKey with the repo and defining the llm: prefix are taken care of automatically in the llm-playground repos. Here we show what you need to do with a repo you create and name.)
PREFIX llm: <http://franz.com/ns/allegrograph/8.0.0/llm/>
INSERT {
?node rdfs:label ?name.
?node rdf:type <http://franz.com/HistoricalFigure>.
} WHERE {
(?name ?node) llm:response "List 50 Historical Figures".
} View the triples. We got 100 (you may get a different number). Here are a bunch:
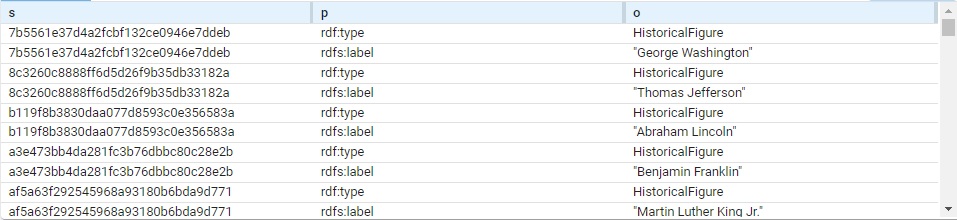
The subject of each triple is a system-created node name. (These are actual values, not blank nodes.) Each is of type HistoricalFigure and each has a label with the actual name.
This query will just list the names (the order is different from above):
# View triples
SELECT ?name WHERE
{ ?node rdf:type <http://franz.com/HistoricalFigure> .
?node rdfs:label ?name . }
name
"Aristotle"
"Plato"
"Socrates"
"Homer"
"Marco Polo"
"Confucius"
"Genghis Khan"
"Cleopatra"
...
(Note: this come from an chat program external to Franz Inc. The choice and morality of the figures selected are beyond our control.)
Now we are going to use these triples to find associatione among them. First we must create a vector database. Create a file historicalFigures.def that looks like this (inserting your aiApi key where indicated):
gpt
openai-api-key "[your aiApi key goes here]"
vector-database-name "historicalFigures"
vector-database-dim 1536
limit 1000000
splitter list
include-predicates <http://www.w3.org/2000/01/rdf-schema\#label> Note the # mark in the last line is escaped. The file in Lineparse format so a # is a comment character unless escaped.
Pass this file to the agtool llm index command like this (filling in the username, password, host, and port number):
% agtool llm index USER:PW@HOST:PORT/historicalFigures historicalFigures.def This gives a fair amount of output (one line per figure and some extra lines). You can redirect the output to a file if you wish, as we show in the example below.
This writes two Vector Database files in the [agdir]/data/rootcatalog/historicalFigures/ directory:
historicalFigures.vdb.dat
historicalFigures.vdb.vec Now we can do more with our data, such as see who are nearest neighbours with this query with a score of at least .5:
PREFIX llm: <http://franz.com/ns/allegrograph/8.0.0/llm/>
PREFIX o: <http://franz.com/>
INSERT {?xnode o:similarTo ?ynode.} WHERE {
?xnode rdfs:label ?xname ;
rdf:type o:HistoricalFigure .
(?ynode ?score ?yname) llm:nearestNeighbor (?xname "historicalFigures" 10 0.5)
FILTER(?xnode != ?ynode)
} Note that we define a o: prefix to avoid having to enter the full URIs, and we use the trick of separating lines with the same subject with a semicolon (and then leaving the subject out).
We used Gruff (click on Gruff to the left) to display a selection of the results (we suppressed the display of the node values):
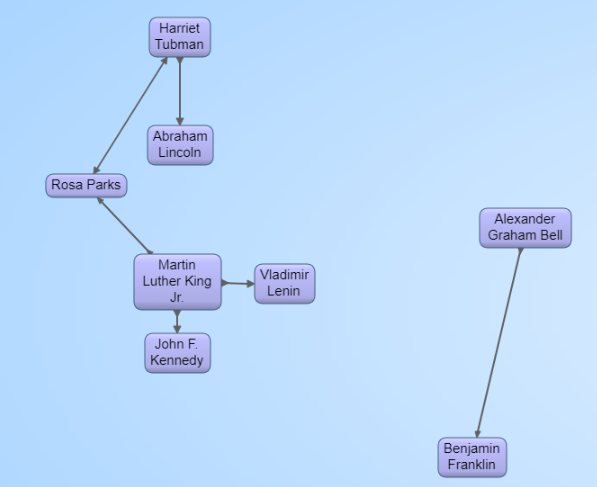
We only obtained 50 historical figures so the number of connections is limited (but the example did run quickly). Do this with several hundred famous people from history and you will see a much richer connections graph.
Retrieval Augmented Generation
In this quite large example, we illustrate retrieval augmented generation (RAG). We start with a database of statements by the linguistics scholar and political commentator Noam Chomsky. We pick him because his writings are easily available and he discusses many significant topics (we take no position on his conclusions).
We start with a collection of triples with initial information. Create a repo name chomsky47. The data for this repo is in a file named chomsky47.nq. This is a large file with over a quarter of a million statements. Its size is about 47MB. Download it by clicking on this link: https://s3.amazonaws.com/franz.com/allegrograph/chomsky47.nq
Once downloaded, load chomsky47.nq into your new chomsky47 AllegroGraph repo.
Set the Query Option for the chomsky47 repo so the openaiApiKey is associated with the repo by going to Repository | Repository Control | Query execution options (type "Query" into the search box after Repository control is displayed) and enter the label openaiApiKey and your key as the value, then click SAVE QUERY OPTIONS (this same action was done before with another repo):

(The key in the picture is not valid.) You can also skip this and use this PREFIX with every query (using your valid key, of course):
PREFIX franzOption_openaiApiKey: <franz:sk-U01ABc2defGHIJKlmnOpQ3RstvVWxyZABcD4eFG5jiJKlmno> Create the file chomsky.def with the following contents, putting your your Openai API key where it says [put your valid AI API key here]
gpt
openai-api-key "[put your valid AI API key here]"
vector-database-name "chomsky47"
vector-database-dim 1536
limit 1000000
splitter list
include-predicates <http://chomsky.com/hasContent> Once the file is ready, run this command supplying your username, password, host, port, and the chomsky.def file path. Note we use 2> redirection which, in bash or bash-comptible shells, causes the output goes to the /tmp/chout file -- there is a great deal of output.
% agtool llm index USER:PASS@HOST:PORT/chomsky47 [LOCATION/]chomsky.def 2> /tmp/chout Try this query:
PREFIX llm: <http://franz.com/ns/allegrograph/8.0.0/llm/>
SELECT ?response ?citation ?score ?content {
# What is your name?
# Is racism a problem in America?
# Do you believe in a 'deep state'?
# What policy would you suggest to reduce inequality in America?
# Briefly explain what you mean by 'Universal Grammar'.
# What is the risk of accidental nuclear war?
# Are you a liberal or a conservative?
# Last time I voted, I was unhappy with both candidates. So I picked the lesser of two evils. Was I wrong to do so?
# How do I bake chocolate chip cookies? (Note: this query should return NO answer.)
bind("What is your name?" as ?query)
(?response ?score ?citation ?content) llm:askMyDocuments (?query "chomsky47" 10 0.0).
} This produces this response:
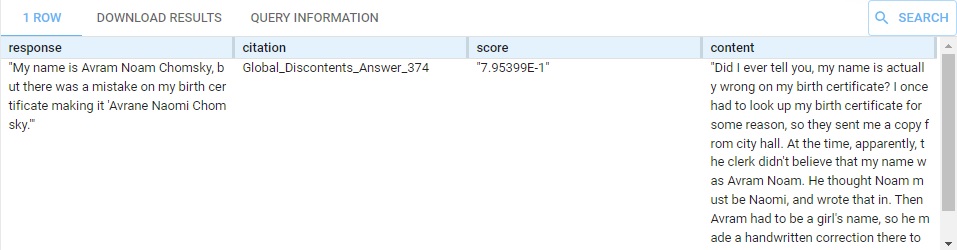
We have (commented out) other questions, about linguistics, politics, and himself (and about cookie recipes). Replace What is your name? with some of those. The content that supports the result is supplied, on multiple rows if it comes from various places (the summary is the same in each row, just the supporting content changes).
serpAPI example
Is this example we use the SERP API for our query. SERP stands for Search Engine Results Page. See https://serpapi.com/ for more information and to obtain a key needed to use the API. Note the openaiApiKey used in earlier queries will not work and you must add this prefix to any SERP API query (replacing the invalid key with a valid one):
PREFIX franzOption_serpApiKey: <franz:11111111111120b7c4d3d04a9837e3362edb2733318effec25c447445dfbf594> See llm for more information on keys are query prefixes.
Here is the query. Paste it into the Query field in AGWebView:
PREFIX llm: <http://franz.com/ns/allegrograph/8.0.0/llm/>
PREFIX franzOption_openaiApiKey: <franz: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>
PREFIX franzOption_serpApiKey: <franz:cb61b7cb2077c507583ab621e57d3d6b04361b7638c76b7959408e17038e161c>
# "Predict tomorrow's top headline baed on today's headlines: "
SELECT (llm:response(concat("Predict tomorrow's top headline based on today's headlines: ", GROUP_CONCAT(?news; separator="; "))) as ?prediction)
{
?news llm:askSerp ("Top news headlines" 10 "top_stories")
}